LOC101059915
In-game article clicks load inline without leaving the challenge.
LOC101059915 is a protein, which in humans is encoded by the LOC101059915 gene. It is located on the X chromosome and has restricted expression in the testis.
Gene
The LOC101059915 gene has two aliases known as chromosome X open reading frame 49-like and BX276092.6.
Locus and Structure
LOC101059915 is located on the X chromosome at locus Xq13.1. It is 1,831 base pairs long and the gene sequence has 6 exons. LOC101059915 also has one protein coding transcript.
Promoter Region and Expression
The promoter region of LOC101059915 is located on the sense strand of DNA, and between base pair 71666098 and 71667904 on the X chromosome. It spans up to 1.806 bp. Expression of LOC101059915, however, is relatively low in human cells, and is primarily limited to the testis.
Protein
General Features and Compositional Analysis
The protein has 518 amino acids and a molecular mass of 55.1 kDa. The isoelectric point is 8.15. Compared to other human proteins LOC101059915 is glycine-, proline-rich, and serine-rich but the protein has lower levels of tyrosine.
Domains
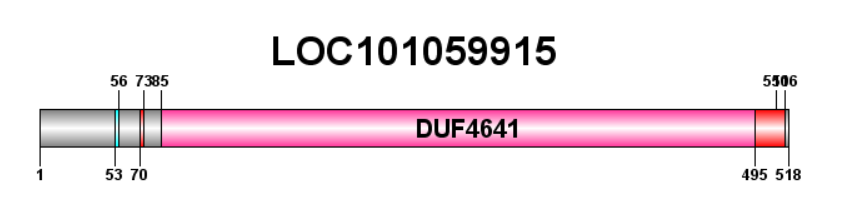
The domain of unknown function, DUF4641 covers almost the entire protein. It is a part of pfam15483. It is 410 amino acids long, from amino acid 85 until amino acid number 495.
Secondary Structure
The secondary structure of LOC101059915 has been shown to consist of primarily alpha helices as determined by models made on I-TASSER and analysis using ExPASy tools.

Post-translation modifications
LOC101055915 is predicted to contain many different post-translational modifications. This include sites for phosphorylation (NetPhos 2.0) and sumoylation (SUMOplot Analysis Program).
Subcellular localization
The LOC101059915 protein has been predicted to be located in the cell nucleus (PSORT II).
Homology and Evolution
Paralogs
CXorf49 and CXorf49B are paralogs of LOC101059915. They share upwards of 78% similarity with LOC101059915 and likely went through a gene duplication event relatively recently, in evolutionary terms, due to the high degree of conservation between all three sequences.
CXorf49 is especially interesting due it being shown to be involved as one of the components of a small group of the HL-60 cell proteome that are most prone to form 4-Hydroxy-2-nonenal(HNE) adducts, upon exposure to nontoxic (10 μM) HNE concentrations, along with heat shock 60 kDa protein 1.
Orthologs
Using BLAST no orthologs for LOC101059915 are found in single celled organisms, fungi or plants whose genomes have been sequenced. For multi-cellular organisms, orthologs are found in mammals, excluding Monotremes. The table below shows a representative sample of 20 of the orthologs for LOC101059915. The table is organized based on the time of divergence from humans in millions of years (MYA). In cases where the divergence time-frame is the same the orthologs are sorted by identity (%).
| Genus and species name | Common name | Divergence from Human Lineage (MYA) | Accession number | Sequence length (aa) | Sequence identity to human protein | Sequence similarity to Human Protein |
|---|---|---|---|---|---|---|
| Pan paniscus | Chimpanzee | 6.65 | XP_003820175.1 | 518 | 97% | 97% |
| Papio anubis | Olive Baboon | 29.4 | XP_003919408.1 | 514 | 70 % | 77% |
| Piliocolobus tephrosceles | Ugandan Red Colobus | 29.4 | XP_023050488.1 | 514 | 68 % | 76% |
| Cebus capucinus imitator | White-Headed Capuchin | 43 | XP_017372264.1 | 472 | 68 % | 72% |
| Callithrix jacchus | Common Marmoset | 43 | XP_008987720.1 | 474 | 67% | 72% |
| Aotus nancymaae | Nancy Ma's Night Monkey | 43 | XP_010822786.1 | 432 | 63% | 66% |
| Saimiri boliviensis boliviensis | Black-capped Squirrel Monkey | 43 | XP_003944512.1 | 521 | 58% | 68% |
| Rattus norvegicus | Brown Rat | 90 | XP_008756683.2 | 136 | 46% | 55% |
| Ictidomys tridecemlineatus | Thirteen-Lined Ground Squirrel | 90 | XP_021576972.1 | 565 | 45% | 57% |
| Canis lupus familiaris | Dog | 96 | XP_850392.2 | 526 | 52% | 64% |
| Panthera pardus | Leopard | 96 | XP_019284263.1 | 472 | 47% | 58% |
| Enhydra lutris kenyoni | Sea Otter | 96 | XP_022348355.1 | 495 | 47% | 59% |
| Ovis aries musimon | Mouflon | 96 | XP_01208848.1 | 539 | 45% | 56% |
| Capra hircus | Goat | 96 | XP_017899210.1 | 538 | 44% | 55% |
| Pantholops hodgsonii | Tibetan Antelope | 96 | XP_005965061.1 | 533 | 44% | 55% |
| Myotis lucifugus | Little brown bat | 96 | XP_006083036.1 | 500 | 42 % | 55% |
| Trichechus manatus latirostris | Florida manatee | 105 | XP_012415455.1 | 505 | 44% | 55% |
| Orycteropus afer afer | Aardvark | 105 | XP_007957133.1 | 477 | 39% | 50% |
| Phascolarctos cinereus | Koala | 159 | XP_020834608.1 | 474 | 37% | 50% |
| Sarcophilus harrisii | Tasmanian Devil | 159 | XP_023362890.1 | 576 | 33% | 50% |
Phylogeny

The most distant ortholog for LOC101059915 is from the species Sarcrophilus harrisii which is commonly known as the Tasmanian Devil dating from more than 159.0 million years ago.